|
|
||||
|
Published by : PROFESSIONAL MEDICAL PUBLICATIONS |
||||
|
ISSN 1681-715X |
||||
|
||||
|
- |
||||
|
ORIGINAL ARTICLE |
||||
|
- |
||||
|
Volume 24 |
April - June 2008 (Part-I) |
Number 2 |
||
|
|
||||
|
|
||||
|
|
||||
|
Published by : PROFESSIONAL MEDICAL PUBLICATIONS |
||||
|
ISSN 1681-715X |
||||
|
||||
|
- |
||||
|
ORIGINAL ARTICLE |
||||
|
- |
||||
|
Volume 24 |
April - June 2008 (Part-I) |
Number 2 |
||
|
|
||||
|
|
||||
Enterotoxigenecity of s. aureus isolates recovered
from chronic urogenital tract infection
in North Palestine
Ghaleb Adwan1, Bassam Abu-Shanab2, Kamel Adwan3, Marwan Odeh4
ABSTRACT
Objectives: Staphylococcus aureus is an important pathogen associated with diseases in a variety of hosts including humans. It produces several toxins and virulence factors that contribute to its pathogenic potential such as staphylococcal enterotoxins (SEs). This study was conducted to determine enterotoxigenicity of S. aureus associated with chronic urogenital tract infection by detecting enterotoxin genes.
Setting: This study was done in The Microbiology laboratory, An-Najah N. University, Palestine.
Methodology: A total of 90 S. aureus isolates recovered from clinical samples from patients suffering from chronic urogenital tract infection in the North of Palestine were used to detect the presence of staphylococcal enterotoxin genes sea, seb, sec, sed and see by polymerase chain reaction (PCR) assay.
Results: Out of 90 S. aureus isolates tested, it was found that 57 (63.3%) of these isolates harboured one or more enterotoxin genes. Up to 78.9% of the enterotoxigenic isolates possessed one SE gene. The majority of these enterotoxigenic strains (61.4%) isolated from both semen and urine samples harbored sec gene either alone or in combination with other genes. Also the prevalence of genes in combination was significantly more common in S. aureus isolates derived from urine 9/33 (27.3%), as compared to those derived from semen 3/24 (12.5%).
Conclusions: The role of enterotoxin genes in the pathogenesis of urogenital tract infection is still unknown. However, it is evident that urogenital infection can be caused by S. aureus strains which lack these genes. Other newly detected genes may play a role in pathogenesis.
KEYWORD:
Enterotoxigenic S. aureus, Staphylococcal enterotoxins, SEs, Palestine.Pak J Med Sci April - June 2008 (Part-I) Vol. 24 No. 2 246-250
1. Ghaleb Adwan,
2. Bassam Abu-Shanab,
Faculty of Veterinary Medicine,
3. Kamel Adwan,
1&3:Department of Biology and Biotechnology,
1-3: An-Najah N. University,
Nablus, Palestine
4. Marwan Odeh,
Al-Hikma Medical laboratories,
Toulkarem, Palestine
Correspondence
Ghaleb Adwan,
Department of Biology and Biotechnology,
An-Najah N. University,
P. O. Box (7) - Nablus, Palestine.
E-mail: adwang@najah.edu
* Received for Publication: August 23, 2007
* Revision Received: August 25, 2007
* Revision Accepted: January 15, 2008
INTRODUCTION
S. aureus
is a commensal bacterium found on host skin and respiratory tract mucosal surfaces. However, it is also associated with diseases in variety of hosts including humans. It is a significant pathogen involved in nosocomial and community-acquired infections in humans. The increasing prevalence of multidrug-resistant S. aureus worldwide is an additional problems and resistance to antimicrobial compounds reduces their effectiveness and increases morbidity, mortality and health care costs worldwide.1 A variety of toxins and virulence factors produced by certain clinical isolates such as hemolysins, exfoliative toxin, and pyrogenic toxin superantigens including enterotoxins and toxic shock syndrome toxin. These toxins are involved in the pathogenesis of staphylococcal diseases such as food poisoning, scalded skin syndrome, and toxic shock syndrome.2 Traditionally, five classic antigenic SE types were recognised SEA to SEE encoded by sea to see genes. During the 1990s, new SE’s SEG to SEJ were reported and their genes described. Recently, data resulting from partial or complete genome sequence analyses have led to a rapid description of further new genes sek, ser and seu.3 The role of several recently described SEs as causative agents in toxin-mediated staphylococcal diseases or in enhancing the severity of S. aureus infections has not been confirmed.4Previously, we have reported the prevalence of enterotoxigenic staphylococci from different clinical human isolates from Northern Palestine, as 41.2%. The majority of these enterotoxigenic isolates carried sea gene (42.9%), and none of positive isolates harboured more than one toxin gene.
5 Also limited information is available on enterotoxigenic S. aureus isolated from urogenital tract infection. Hence this study was conducted to determine the prevalence of enterotoxigenic S. aureus isolates associated with chronic urogenital tract infection by detecting sea, seb, sec, sed and see genes.METHODOLOGY
During a one-year-period between August of 2005 and July of 2006, a total of 90 clinical isolates of S. aureus (51 derived from urine samples and 39 from semen) were recovered from patients suffering from chronic urogenital tract infection. These isolates tested to detect the presence of SE genes sea, seb, sec, sed and see by PCR assay. All isolates were identified in the microbiology laboratories of An-Najah National University, Palestine, by Gram stain, culture properties on nutrient agar and mannitol salt agar, detection of hemolysis on 5% sheep blood agar and by tube coagulase reaction.
Methicillin resistance was carried out using the disk diffusion method.
6 Methicillin (5µg) disks (Oxoid) were used, inhibition zones were determined in accordance with procedures of the National Committee for Clinical Laboratory Standard.7 Isolates giving an inhibition zone >14mm were confirmed as methicillin-sensitive isolate, while others were considered as resistant.DNA template was isolated from approximately 10 colonies of the bacteria as described previously.
5 Bacterial cells were washed once with 1.0ml of 0.02 M sodium phosphate (Na2HPO4.2H2O) pH 7.4 in 0.9% NaCl and centrifugation at 12000Xg for ten minutes. The pellet was then resuspended in 200µl of lysis buffer (1mM EDTA, 10mM Tris-chloride, pH 8) with 12U lysostaphin (Sigma) and incubated for 60 minutes at 37°C. Then 4.5 U of proteinase K (MO BIO) were added and incubated for 45 minutes at 60°C, and not for 10 minutes at 95°C. The total DNA is spun at 12,000 X g for 15 seconds and stored at -20°C for DNA amplification. The sea, seb, sec, sed and see gene sequences were detected using the primer pairs described previously.8For PCR amplification, the total reaction mixture was 30µl, and performed as previously described.
5,9 It included 10pmol of each primer, 0.2mM of each deoxynucleoside triphosphate (PeQLab), 1 X PCR reaction buffer (PeQLab), 1.5mM of MgCl2 (PeQLab), 1U of Taq DNA polymerase (PeQLab). Finally, 1µl of DNA template was added to each 0.2-ml reaction tube. The amplification was carried out using the thermal cycler (Mastercycler personal, Eppendorf) with the following program: 1 X four minutes precycle at 94°C followed by 30 PCR cycles (2 minutes at 94°C, 2 minutes at 55°C, and one minute at 72°C). At the end of the cycles, the reaction mixture was maintained at 72°C for five minutes. PCR products were separated by electrophoresis in 2% agarose gel and stained with ethidium bromide. Positive reference strains for these genes were also included. All primers used in this study were synthesized by Integrated DNA Technologies (IDT), Inc. USA.RESULTS
Ninety isolates of clinical S. aureus from chronic urogenital tract infection were studied for methicillin resistance. It was found that fifty two (63.4%) and five out of eight methicillin sensitive isolates (62.5%) of methicillin resistant and methicillin sensitive isolates, respectively, were enterotoxigenic (Table-I).

Out of 90 S. aureus isolates tested for SE genes, it was found that 57 (63.3%) of these detected isolates harboured one or more enterotoxin genes. Up to 78.9% of the enterotoxigenic strains carried one SE gene. The majority of these positive toxin gene isolates 35/57 (61.4%), were sec-positive either single or combined with other genes.
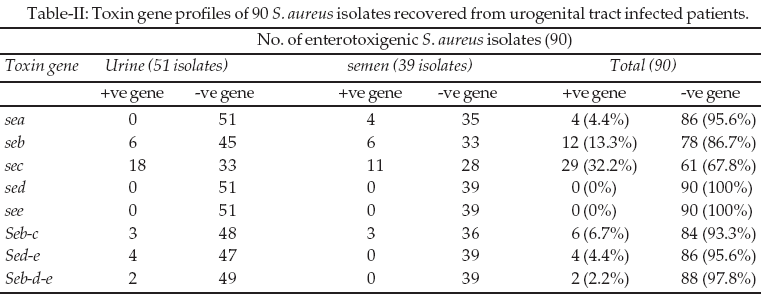
Toxin gene profiles of these 90 enterotoxigenic S. aureus isolates are presented in Table-II. It also found that gene combination was significantly more common in S. aureus isolates derived from urine 9/33 (27.3%), while in S. aureus isolates derived from semen 3/24 (12.5%).
DISCUSSION
The literature shows variable results concerning the occurrence of enterotoxigenic S. aureus strains in clinical samples. This is probably due to the differences among the origin of clinical samples, number of test samples, sensitivity of methods, and types of enterotoxins or enterotoxin genes that were detected. Indeed prevalence of enterotoxigenic S. aureus observed in different human clinical samples varies markedly between reports.
5,10-14 Only 63.3% of the our isolates from patients suffering from chronic urogenital infection in the from North of Palestine were enterotoxigenic, This study showed higher prevalence than our previous report,5 which was 41.2% of test isolates. If other more recently described toxin genes in S. aureus were taken into account the prevalence might be higher than being reported.It is important to mention that the detection of the toxin genes by genotypic techniques like PCR in Palestinian S. aureus isolates does not necessarily indicate the ability of these isolates to produce biologically active toxin or sufficient level of toxin to induce disease. To confirm the toxicity of these strains it is important to demonstrate the toxin levels that are produced by these enterotoxigenic strains.
13 The presence of a higher number of leukocytes in urine or semen samples in case of urogenital staphylococcal infection could be a factor that contributes genetic selection of strains that can produce superantigens. These toxins, others may have a damaging effect on that cell type. Those urogenital strains which carried enterotoxin genes may induce releasing of specific cytokines that may inhibit the efficiency of human immune response by their expressions; this may contribute to the persistence of S. aureus in urogenital tract and promote inflammation in these tissues or enhance the chronicity of this disease.In the present study, the majority of the enterotoxigenic S. aureus isolates carried sec gene alone or in combination with other genes. These results were consistent with previous reports which showed that most human clinical enterotoxigenic S. aureus isolates harbored sec and seb genes.
15,16 However, these results were in contrast to other studies which showed that most human clinical enterotoxigenic S. aureus isolates carried seg and sei genes,17,18 seb gene,19,20 or sea gene.5,10,11,21 This discrepancy in prevalence is possibly due to geographical differences, which may be related to different ecological origins of the isolated strains and the genes which have been detected. Our results in this study are different in many respects from our report that was published previously,5 which has showed that all enterotoxigenic isolates detected previously harboured only one toxin gene. The possession of more than a single enterotoxin gene was reported, the multiple genes and their frequencies were variable in these studies.4,10,11,14-16 The presence of multiple expressed toxins in a single S. aureus strain makes it less clear whether any single toxin is responsible for diseases.22 In spite of this, the possession of multiple virulence genes supports a strong potential of the isolates to cause severe illness.The role of sea, seb, sec, sed and see genes in the pathogenesis of urogenital tract infection is still unknown. However it is possible that urogenital infection can be caused by S. aureus strains at least lack these genes. Other newly detected genes may play a role in pathogenesis.
Note: All reagents and equipments used in this work were used in a previous project (reference 10) and that project was completely supported by the USA-AED (Academy for Education Development)
REFERENCES
1. Coast J, Smith RD. Solving the problem of antimicrobial resistance: Is a global approach necessary? Drug Discov Today 2003;8:1-2.
2. Schmidt H, Hensel M. Pathogenicity islands in bacterial pathogenesis. Clin Microbiol Rev 2004;17:14-56.
3. Nakayama A, Okayama A, Hashida M, Yamamoto Y, Takebe H, Ohnaka T, et al. Development of a routine laboratory direct detection system of staphylococcal enterotoxin genes. J Med Microbiol 2006;55:273-7.
4. Becker K, Friedrich AW, Lubritz G, Weilert M, Peters G, Von Eiff C. Prevalence of genes encoding pyrogenic toxin superantigens and exfoliative toxins among strains of Staphylococcus aureus isolated from blood and nasal specimens. J Clin Microbiol 2003;41:1434-9.
5. Adwan GM, Abu-shanab AB, Adwan KM, Jarrar NR. Toxigenicity of Staphylococcus aureus isolates from Northern Palestine. Emirates Med J 2006;24:127-9.
6. Bauer AW, Kirby WMM, Sherris JC, Truck M. Antibiotic susceptibility testing by a standardized single disc method. Am J Clin Pathol 1966;45:493-6.
7. National Committee for Clinical Laboratory Standards (NCCLS). Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals. (M31-A). Pennsylvania, USA; 1999.
8. Akineden O, Annemuller C, Hassan AA, Lammler Ch, Wolter W, Zschock M. Toxin genes and other characteristics of Staphylococcus aureus isolates from milk of cows with mastitis. Clin Diagn Lab Immunol 2000;8:959-64.
9. Adwan G, Abu-shanab B, Adwan K. Enterotoxigenic Staphylococcus aureus in raw milk in the North of Palestine. Turk J Biol 2005;29:229-32.
10. Naffa RG, Bdour SM, Migdadi HM, Shehabi AA. Enterotoxicity and genetic variation among clinical Staphylococcus aureus isolates in Jordan. J Med Microbiol 2006;55:183-7.
11. Efuntoye MO, Adetosoye AI. Enterotoxigenicity and drug sensitivity of staphylococci from children aged five years and below with sporadic diarrhoea. East Afr Med J 2003;80:656-9.
12. Mempel M, Lina G, Hojka M, Schnopp C, Seidl HP, Schafer T. High prevalence of superantigens associated with the egc locus in Staphylococcus aureus isolates from patients with atopic eczema. Eur J Clin Microbiol Infect Dis 2003;22:306-9.
13. Omoe K, Ishikawa M, Shimoda Y, Hu DL, Ueda S, Shinagawa K. Detection of seg, seh, and sei genes in Staphylococcus aureus isolates and determination of the enterotoxin productivities of S. aureus isolates harboring seg, seh, or sei genes. J Clin Microbiol 2002;40:857-62.
14. Fueyo JM, Martin MC, Gonzalez-Hevia MA, Mendoza MC. Enterotoxin production and DNA fingerprinting in Staphylococcus aureus isolated from human and food samples. Relations between genetic types and enterotoxins. Int J Food Microbiol 2001;67:139-45.
15. Adesiyun AA, Lenz W, Schaal KP. Phage susceptibility, enterotoxigenicity and antibiograms of Staphylococcus aureus strains isolated from human wounds and diarrhoea. Zentralbl Bakteriol 1992;277:250-9.
16. Reina J, Salva F, Alomar P. Enterotoxin production by strains of Staphylococcus aureus isolated from clinical specimens. J Clin Pathol 1989;42:217-8.
17. Omoe K, Hu DL, Takahashi-Omoe H, Nakane A, Shinagawa K. Identification and characterization of a new staphylococcal enterotoxin-related putative toxin encoded by two kinds of plasmids. Infect Immun 2003;71:6088-94.
18. El-Huneidi W, Bdour S, Mahasneh A. Detection of enterotoxin genes seg, seh, sei, and sej and of a novel aroA genotype in Jordanian clinical isolates of Staphylococcus aureus. Diagn Microbiol Infect Dis 2006;56:127-32.
19. Fluer FS, Sheveleva AF, Tekut’ev SI. Occurrence of Staphylococcus aureus enterotoxigenic strains isolated from pregnant women with pathology. Zh Mikrobiol Epidemiol Immunobiol 2000;6:13-5. [Abstract in English].
20. El-Ghodban A, Ghenghesh KS, Marialigeti K, Tawil A. Enterotoxins and phage typing of Staphylococcus aureus isolated from clinical material and food in Libya. Arch Inst Pasteur Tunis 1999;76:23-5.
21. Fluer FS, Prokhorov VIa, Bondarenko VM, Dmitrienko OA, Lashenkova NN, Men’shikova ED. Isolation rate of enterotoxigenic staphylococci in patients with sepsis, pneumonia and burns. Zh. Mikrobiol Epidemiol Immunobiol 2005;5:3-6. [Abstract in English]
22. Orwin PM, Leung DY, Donahue HL, Novick RP, Schlievert PM. Biochemical and biological properties of staphylococcal enterotoxin K. Infect Immun.2001;69:360-6.
HOME | SEARCH | CURRENT ISSUE | PAST ISSUES
Professional
Medical Publications
Room No. 522, 5th Floor, Panorama Centre
Building No. 2, P.O. Box 8766, Saddar, Karachi - Pakistan.
Phones : 5688791, 5689285 Fax : 5689860
pjms@pjms.com.pk